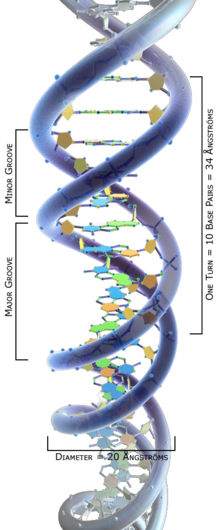
DNA

Deoxyribonucleic acid (DNA) is a nucleic acid that contains the genetic instructions for the development and function of living organisms. All living things contain DNA, with the exception of some viruses with RNA genomes. The main role of DNA in the cell is the long term storage of information. It is often compared to a blueprint, since it contains the instructions to construct other components of the cell, such as proteins and RNA molecules. The DNA segments that carry genetic information are called genes, but other DNA sequences have structural purposes, or are involved in regulating the expression of genetic information. In eukaryotes such as animals and plants, DNA is stored inside the cell nucleus, while in prokaryotes such as bacteria, the DNA is in the cell's cytoplasm. Unlike enzymes, DNA does not act directly on other molecules; rather, various enzymes act on DNA and copy its information into either more DNA, in DNA replication, or transcribe it into protein. In chromosomes, chromatin proteins such as histones compact and organize DNA, as well as helping control its interactions with other proteins in the nucleus. DNA is a long polymer of simple units called nucleotides, which are held together by a backbone made of sugars and phosphate groups. This backbone carries four types of molecules called bases and it is the sequence of these four bases that encodes information. The major function of DNA is to encode the sequence of amino acid residues in proteins, using the genetic code. To read the genetic code, cells make a copy of a stretch of DNA in the nucleic acid RNA. These RNA copies can then be used to direct protein synthesis, but they can also be used directly as parts of ribosomes or spliceosomes.
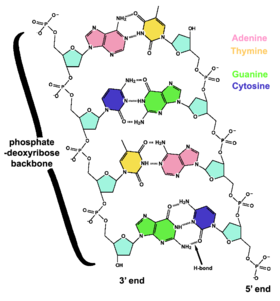
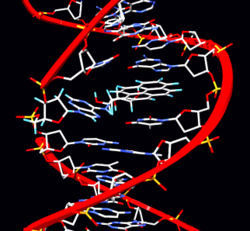
The two strands of DNA are held together by hydrogen bonds between bases. The sugars in the backbone are shown in light blue.
DNA is a long polymer made from repeating units called nucleotides.[1][2] The DNA chain is 22 to 24 angstroms wide and one nucleotide unit is 3.3 angstroms long.[3] Although these repeating units are very small, DNA polymers can be enormous molecules containing millions of nucleotides. For instance, the largest human chromosome is 220 million base pairs long.[4]
In living organisms, DNA does not usually exist as a single molecule, but instead as a tightly-associated pair of molecules.[5][6] These two long strands entwine like vines, in the shape of a double helix. The nucleotide repeats contain both the backbone of the molecule, which holds the chain together, and a base, which interacts with the other DNA strand in the helix. In general, a base linked to a sugar is called a nucleoside and a base linked to a sugar and one or more phosphate groups is called a nucleotide. If multiple nucleotides are linked together, as in DNA, this polymer is referred to as a polynucleotide.[7]
The backbone of the DNA strand is made from alternating phosphate and sugar residues.[8] The sugar in DNA is the pentose (five carbon) sugar 2-deoxyribose. The sugars are joined together by phosphate groups that form phosphodiester bonds between the third and fifth carbon atoms in the sugar rings. These asymmetric bonds mean a strand of DNA has a direction. In a double helix the direction of the nucleotides in one strand is opposite to their direction in the other strand. This arrangement of DNA strands is called antiparallel. The asymmetric ends of a strand of DNA bases are referred to as the 5' (five prime) and 3' (three prime) ends. One of the major differences between DNA and RNA is the sugar, with 2-deoxyribose being replaced by the alternative pentose sugar ribose in RNA.[6]
The DNA double helix is held together by hydrogen bonds between the bases attached to the two strands. The four bases found in DNA are adenine (abbreviated A), cytosine (C), guanine (G) and thymine (T). These four bases are shown below and are attached to the sugar/phosphate to form the complete nucleotide, as shown for adenosine monophosphate.
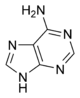
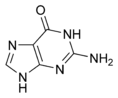
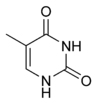
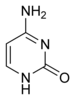
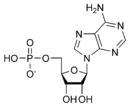
These bases are classified into two types, adenine and guanine are fused five- and six-membered heterocyclic compounds called purines, while cytosine and thymine are six-membered rings called pyrimidines.[7] A fifth pyrimidine base, called uracil (U), replaces thymine in RNA and differs from thymine by lacking a methyl group on its ring. Uracil is normally only found in DNA as a breakdown product of cytosine, but a very rare exception to this rule is a bacterial virus called PBS1 that contains uracil in its DNA.[9]
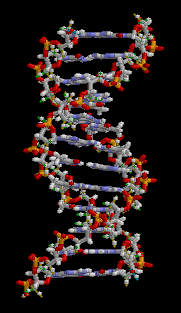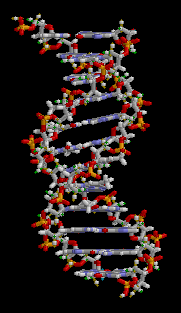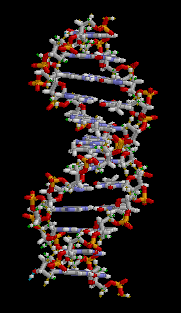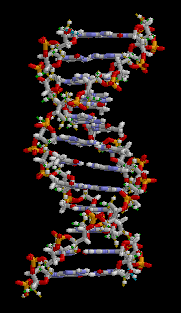
The double helix is a right-handed spiral. As the DNA strands wind around each other, they leave gaps between each set of phosphate backbones, revealing the sides of the bases inside (see animation). There are two of these grooves twisting around the surface of the double helix: one groove is 22 angstroms wide and the other 12 angstroms wide.[11] The larger groove is called the major groove, while the smaller, narrower groove is called the minor groove. The narrowness of the minor groove means that the edges of the bases are more accessible in the major groove. As a result, proteins like transcription factors that can bind to specific sequences in double-stranded DNA usually read the sequence by making contacts to the sides of the bases exposed in the major groove.[12]
 |
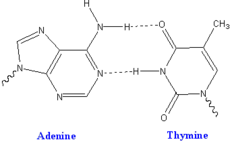 |
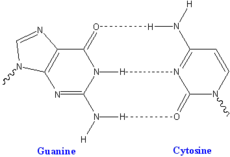
Each type of base on one strand forms a bond with just one type of base on the other strand. This is called complementary base pairing. Here, purines form hydrogen bonds to pyrimidines, with A bonding only to T, and C bonding only to G. This arrangement of two nucleotides joined together across the double helix is called a base pair. In a double helix, the two strands are also held together by forces generated by the hydrophobic effect and pi stacking, but these forces are not affected by the sequence of the DNA.[13] As hydrogen bonds are not covalent, they can be broken and rejoined relatively easily. The two strands of DNA in a double helix can therefore be pulled apart like a zipper, either by a mechanical force or high temperature.[14] As a result of this complementarity, all the information in the double-stranded sequence of a DNA helix is duplicated on each strand, which is vital in DNA replication. Indeed, this reversible and specific interaction between complementary base pairs is critical for all the functions of DNA in living organisms.[1]
The two types of base pairs form different numbers of hydrogen bonds, AT forming two hydrogen bonds, and GC forming three hydrogen bonds (see figures, left). The GC base-pair is therefore stronger than the AT base pair. As a result, it is both the percentage of GC base pairs and the overall length of a DNA double helix that determine the strength of the association between the two strands of DNA. Long DNA helices with a high GC content have strongly interacting strands, while short helices with high AT content have weakly interacting strands.[15] Parts of the DNA double helix that need to separate easily, such as the TATAAT Pribnow box in bacterial promoters, tend to have sequences with a high AT content, making the strands easier to pull apart.[16] In the laboratory, the strength of this interaction can be measured by finding the temperature required to break the hydrogen bonds, their melting temperature (also called Tm value). When all the base pairs in a DNA double helix melt, the strands separate and exist in solution as two entirely independent molecules. These single-stranded DNA molecules have no single shape, but some conformations are more stable than others.[17] The base pairing, or lack of it, can create various topologies at the DNA end. These can be exploited in biotechnology.
DNA is copied into RNA by RNA polymerase enzymes that only work in the 5' to 3' direction.[18] A DNA sequence is called "sense" if its sequence is copied by these enzymes and then translated into protein. The sequence on the opposite strand is complementary to the sense sequence and is therefore called the "antisense" sequence. Both sense and antisense sequences can exist on different parts of the same strand of DNA. In both prokaryotes and eukaryotes, antisense sequences are transcribed, but the functions of these RNAs are not entirely clear.[19] One proposal is that antisense RNAs are involved in regulating gene expression through RNA-RNA base pairing.[20]
A few DNA sequences in prokaryotes and eukaryotes, and more in plasmids and viruses, blur the distinction made above between sense and antisense strands by having overlapping genes.[21] In these cases, some DNA sequences do double duty, encoding one protein when read 5' to 3' along one strand, and a second protein when read in the opposite direction (still 5' to 3') along the other strand. In bacteria, this overlap may be involved in the regulation of gene transcription.[22] While in viruses, overlapping genes increase the amount of information that can be encoded within the small viral genome.[23] Another way of reducing genome size is seen in some viruses that contain linear or circular single-stranded DNA as their genetic material.[24][25]
Benzopyrene, the major mutagen in tobacco smoke, in an adduct to DNA.[48]
DNA can be damaged by many different sorts of mutagens. These include oxidizing agents, alkylating agents and also high-energy electromagnetic radiation such as ultraviolet light and x-rays. The type of DNA damage produced depends on the type of mutagen. For example, UV light mostly damages DNA by producing thymine dimers, which are cross-links between adjacent pyrimidine bases in a DNA strand.[49] On the other hand, oxidants such as free radicals or hydrogen peroxide produce multiple forms of damage, including base modifications, particularly of guanosine, as well as double-strand breaks.[50] It has been estimated that in each human cell, about 500 bases suffer oxidative damage per day.[51][52] Of these oxidative lesions, the most damaging are double-strand breaks, as they can produce point mutations, insertions and deletions from the DNA sequence, as well as chromosomal translocations.[53]
Many mutagens intercalate into the space between two adjacent base pairs. These molecules are mostly polycyclic, aromatic, and planar molecules and include ethidium, proflavin, daunomycin, doxorubicin and thalidomide. DNA intercalators are used in chemotherapy to inhibit DNA replication in rapidly-growing cancer cells.[54] In order for an intercalator to fit between base pairs, the bases must separate, distorting the DNA strand by unwinding of the double helix. These structural modifications inhibit transcription and replication processes, causing both toxicity and mutations. As a result, DNA intercalators are often carcinogens, with benzopyrene diol epoxide, acridines, aflatoxin and ethidium bromide being well-known examples.[55][56]
Overview of biological functions
DNA contains the genetic information that allows living things to function,
grow and reproduce. This information is held in the sequence of pieces of DNA
called genes. Genetic information in genes is transmitted through complementary
base pairing. For example, when a cell uses the information in a gene, the DNA
sequence is copied into a complementary RNA sequence in a process called transcription.
Usually, this RNA copy is then used to make a matching protein sequence in a
process called translation. Alternatively, a cell may simply copy its genetic
information in a process called DNA replication. The details of these functions
are covered in other articles, here we focus on the interactions that happen
in these processes between DNA and other molecules.
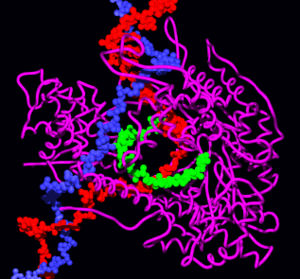
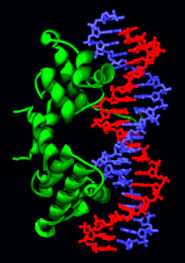
T7 RNA polymerase producing a mRNA (green) from a DNA template (red and blue). The protein is shown as a purple ribbon.[57]
DNA is located in the cell nucleus of eukaryotes, as well as small amounts in mitochondria and chloroplasts. In prokaryotes, the DNA is held within an irregularly shaped body in the cytoplasm called the nucleoid.[59] The DNA is usually in linear chromosomes in eukaryotes, and circular chromosomes in prokaryotes. In the human genome, there is approximately 3 billion base pairs of DNA arranged into 46 chromosomes.[60] The genetic information in a genome is held within genes. A gene is a unit of heredity and is a region of DNA that influences a particular characteristic in an organism. Genes contain an open reading frame that can be transcribed, as well as regulatory sequences such as promoters and enhancers, which control the expression of the open reading frame.
In many species, only a small fraction of the total sequence of the genome encodes protein. For example, only about 1.5% of the human genome consists of protein-coding exons, with over 50% of human DNA consisting of non-coding repetitive sequences.[61] The reasons for the presence of so much non-coding DNA in eukaryotic genomes and the extraordinary differences in genome size, or C-value, among species represent a long-standing puzzle known as the "C-value enigma".[62]
Some non-coding DNA sequences play structural roles in chromosomes. Telomeres and centromeres typically contain few genes, but are important for the function and stability of chromosomes.[38][63] An abundant form of non-coding DNA in humans are pseudogenes, which are copies of genes that have been disabled by mutation.[64] These sequences are usually just molecular fossils, although they can occasionally serve as raw genetic material for the creation of new genes through the process of gene duplication and divergence.[65]
Interactions with proteins
All the functions of DNA depend on interactions with proteins. These protein
interactions can either be non-specific, or the protein can only bind to a particular
DNA sequence. Enzymes can also bind to DNA and of these, the polymerases that
copy the DNA base sequence in transcription and DNA replication are particularly
important.
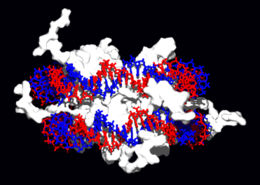 |
Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes between DNA and structural proteins. These proteins organize the DNA into a compact structure called chromatin. In eukaryotes this structure involves DNA binding to a complex of small basic proteins called histones, while in prokaryotes multiple types of proteins are involved.[66] The histones form a disk-shaped complex called a nucleosome, which contains two complete turns of double-stranded DNA wrapped around its surface. These non-specific interactions are formed through basic residues in the histones making ionic bonds to the acidic sugar-phosphate backbone of the DNA, and are therefore largely independent of the base sequence.[67] Chemical modifications of these basic amino acid residues include methylation, phosphorylation and acetylation.[68] These chemical changes alter the strength of the interaction between the DNA and the histones, making the DNA more or less accessible to transcription factors and changing the rate of transcription.[69] Other non-specific DNA-binding proteins found in chromatin include the high-mobility group proteins, which bind preferentially to bent or distorted DNA.[70] These proteins are important in bending arrays of nucleosomes and arranging them into more complex chromatin structures.[71]
A distinct group of DNA-binding proteins are the single-stranded DNA-binding
proteins. that specifically bind single-stranded DNA. In humans, replication
protein A is the best-characterised member of this family and is essential for
most processes where the double helix is separated, including DNA replication,
recombination and DNA repair.[72] These binding proteins seem to stabilize single-stranded
DNA and protect it from forming stem loops or being degraded by nucleases.
The lambda repressor helix-turn-helix transcription factor bound to its DNA target.[73]
In contrast, other proteins have evolved to specifically bind particular DNA sequences. The most intensively studied of these are the various classes of transcription factors. These proteins control gene transcription. Each one of these proteins bind to one particular set of DNA sequences and thereby activates or inhibits the transcription of genes with these sequences close to their promoters. The transcription factors do this in two ways. Firstly, they can bind the RNA polymerase responsible for transcription, either directly or through other mediator proteins, this locates the polymerase at the promoter and allows it to begin transcription.[74] Alternatively, transcription factors can bind enzymes that modify the histones at the promoter, this will change the accessibility of the DNA template to the polymerase.[75]
As these DNA targets can occur throughout an organism's genome, changes in
the activity of one type of transcription factor can affect thousands of genes.[76]
Consequently, these proteins are often the targets of the signal transduction
processes that mediate responses to environmental changes or cellular differentiation
and development. The specificity of these transcription factors' interactions
with DNA come from the proteins making multiple contacts to the edges of the
DNA bases, allowing them to "read" the DNA sequence. Most of these
base interactions are made in the major groove, where the bases are most accessible.[77]
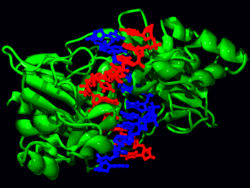
The restriction enzyme EcoRV (green) in a complex with its substrate DNA.[78]
Forensics
Forensic scientists can use DNA in blood, semen, skin, saliva or hair at a crime scene to identify a perpetrator. This process is called genetic fingerprinting or more accurately, DNA profiling. In DNA profiling, the lengths of variable sections of repetitive DNA, such as short tandem repeats and minisatellites, are compared between people. This method is usually an extremely reliable technique for identifying a criminal.[93] However, identification can be complicated if the scene is contaminated with DNA from several people.[94] DNA profiling was developed in 1984 by British geneticist Sir Alec Jeffreys,[95] and first used in forensic science to convict Colin Pitchfork in the 1988 Enderby murders case.[96] People convicted of certain types of crimes may be required to provide a sample of DNA for a database. This has helped investigators solve old cases where only a DNA sample was obtained from the scene. DNA profiling can also be used to identify victims of mass casualty incidents.[97]
Bioinformatics
Bioinformatics involves the manipulation, searching, and data mining DNA sequence
data. The development of techniques to store and search DNA sequences have led
to widely-applied advances in computer science, especially string searching
algorithms, machine learning and database theory.[98] String searching or matching
algorithms, which find an occurrence of a sequence of letters inside a larger
sequence of letters, was developed to search for specific sequences of nucleotides.[99]
In other applications such as text editors, even simple algorithms for this
problem usually suffice, but DNA sequences cause these algorithms to exhibit
near-worst-case behaviour due to their small number of distinct characters.
The related problem of sequence alignment aims to identify homologous sequences
and locate the specific mutations that make them distinct. These techniques,
especially multiple sequence alignment, are used in studying phylogenetic relationships
and protein function.[100] Data sets representing entire genomes' worth of DNA
sequences, such as those produced by the Human Genome Project, are difficult
to use without annotations, which label the locations of genes and regulatory
elements on each chromosome. Regions of DNA sequence that have the characteristic
patterns associated with protein- or RNA-coding genes can be identified by gene
finding algorithms, which allow researchers to predict the presence of particular
gene products in an organism even before they have been isolated experimentally.[101]
DNA was first isolated by Friedrich Miescher who discovered a substance he called "nuclein" in 1869.[109] In 1929 this discovery was followed by Phoebus Levene's identification of the base, sugar and phosphate nucleotide unit.[110] Levene suggested that DNA consisted of a string of nucleotide units linked together through the phosphate groups. However Levene thought the chain was short and the bases repeated in a fixed order. In 1937 William Astbury produced the first X-ray diffraction patterns that showed that DNA had a regular structure.[111]
In 1943, Oswald Theodore Avery discovered that traits of the "smooth" form of the Pneumococcus could be transferred to the "rough" form of the same bacteria by mixing killed "smooth" bacteria with the live "rough" form. Avery identified DNA as this transforming principle.[112] DNA's role in heredity was confirmed in 1953, when Alfred Hershey and Martha Chase in the Hershey-Chase experiment, showed that DNA is is the genetic material of the T2 phage.[113]
Using X-ray diffraction data from Rosalind Franklin and the information that the bases were paired, James D. Watson and Francis Crick produced the first accurate model of DNA structure in 1953 in their article The Molecular structure of Nucleic Acids.[5] Watson and Crick proposed the central dogma of molecular biology in 1957, describing how proteins are produced from nucleic DNA. In 1962 Watson, Crick, and Maurice Wilkins jointly received the Nobel Prize.[114]
In an influential presentation in 1957, Crick laid out the "Central Dogma",
which foretold the relationship between DNA, RNA, and proteins, and articulated
the "adaptor hypothesis".[115] Final confirmation of the replication
mechanism that was implied by the double-helical structure followed in 1958
through the Meselson-Stahl experiment.[116] Further work by Crick and coworkers
showed that the genetic code was based on non-overlapping triplets of bases,
called codons, allowing Har Gobind Khorana, Robert W. Holley and Marshall Warren
Nirenberg to decipher the genetic code.[117] These findings represent the birth
of molecular biology.
We also recommend the excellent DNA tutorials at Cold Spring Harbour